| Tools |
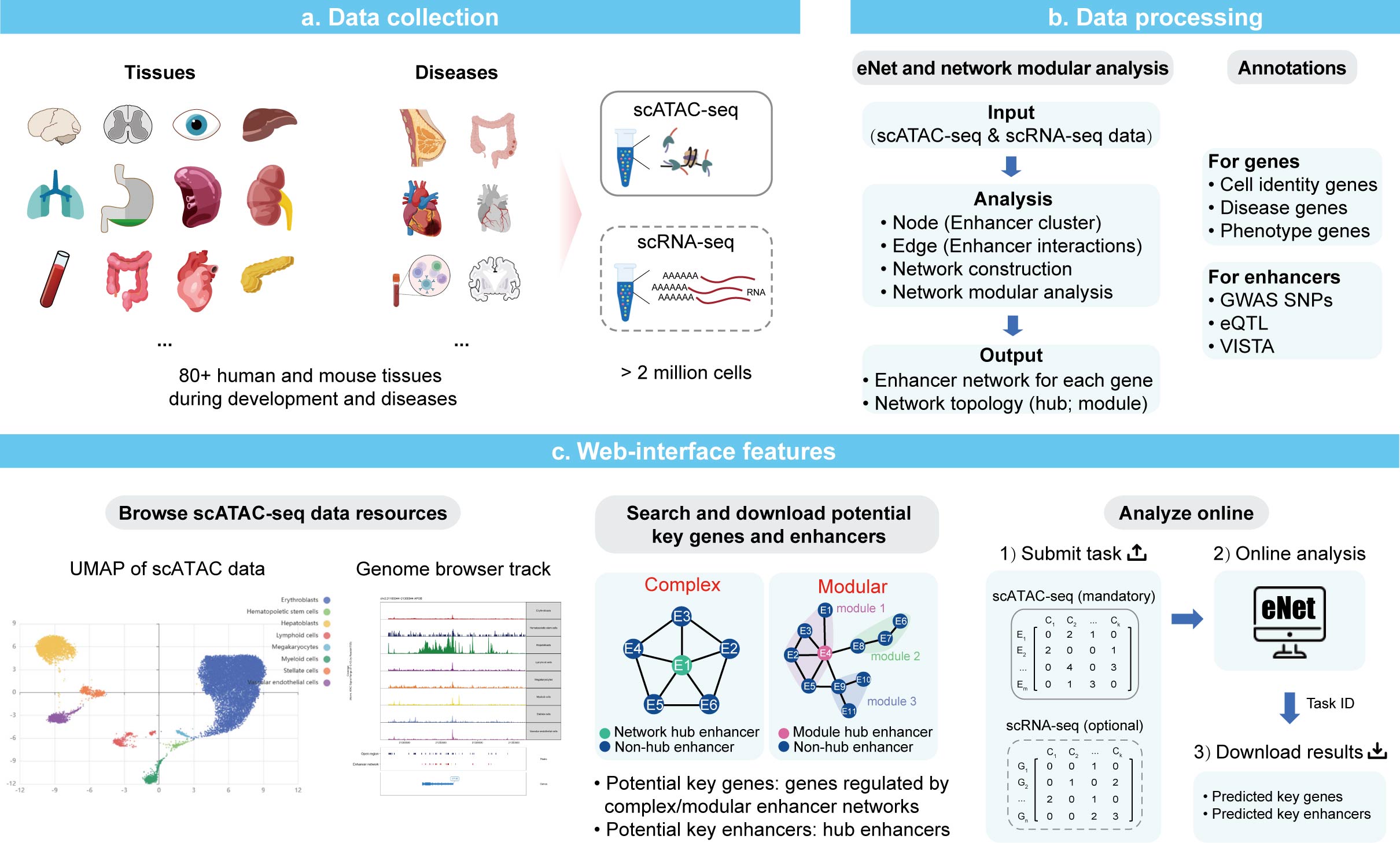
Repositories Webserver eNetDB is database provide potential key genes and enhancers under kinds of biological systems for experimental validation, covering 80+ tissues/cell types that include a large number of healthy/diseased tissues of human and mouse.
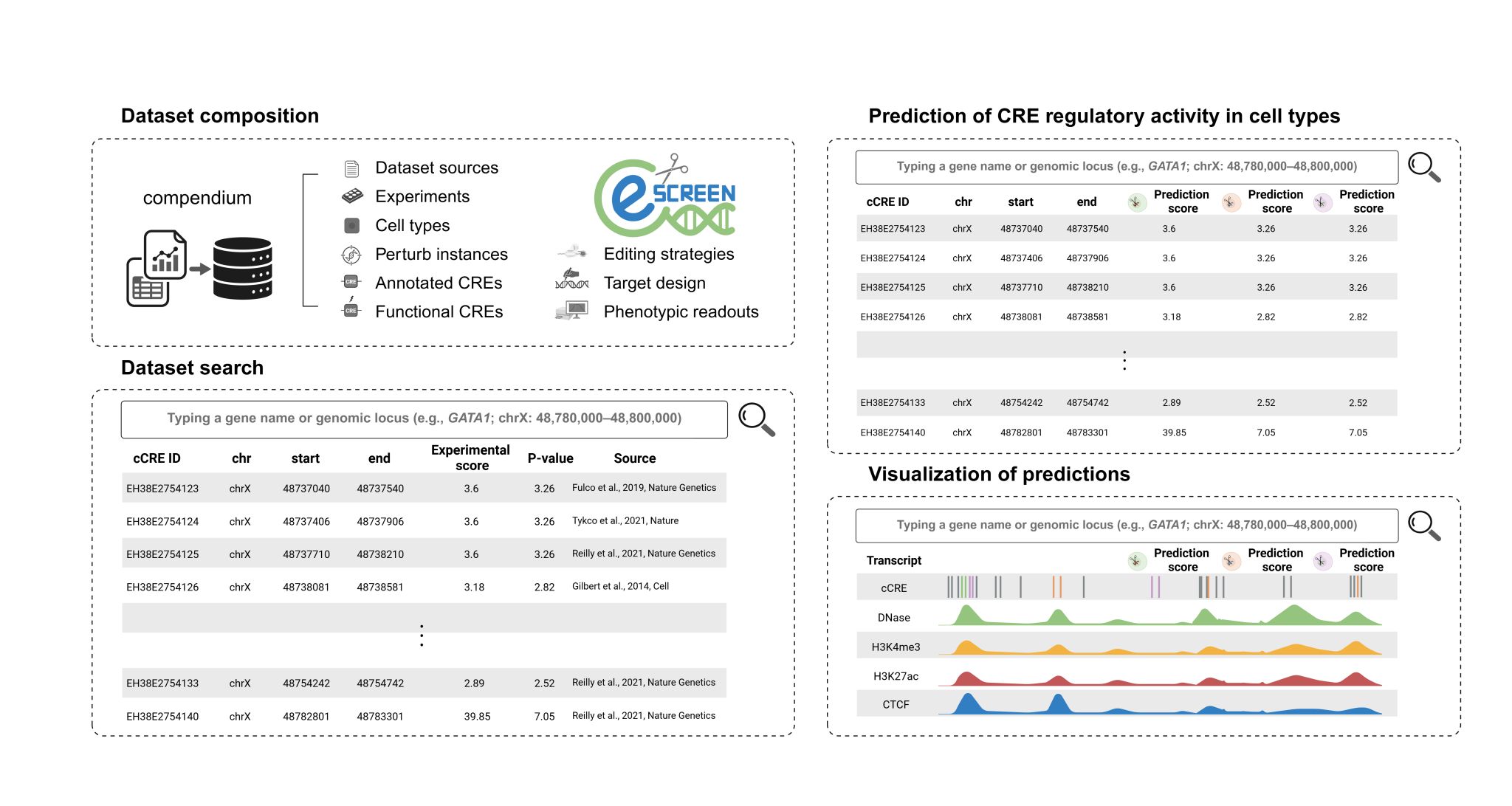
eScreen is a sequence-sensitive model built upon the Striped Hyena2 architecture designed to learn interpretable regulatory context model from CRISPR perturbation experiment. Using the results of CRISPR perturbation experiment analysis and information about transcriptional factor motif, eScreen learns functional regulatory syntax and predicts regulatory activity of cis-regulatory elements.
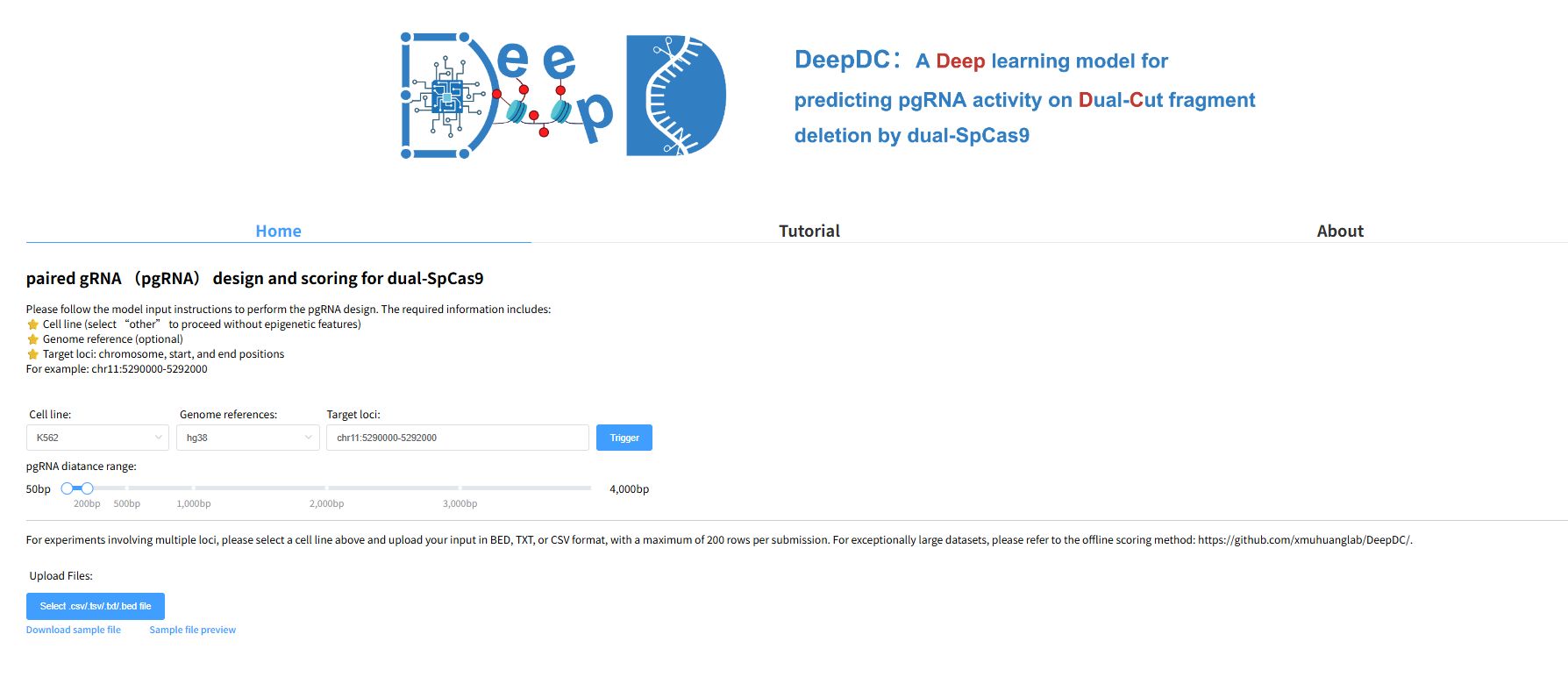
DeepDC, developed by Jialiang Huang's Lab at Xiamen University (XMU) and Teng Fei's Lab at Northeastern University (NEU), is a deep learning model for predicting pgRNA activity on dual-cut fragment deletion by dual-SpCas9. It combines a HyenaDNA block and XGBoost to extract epigenomic features and sequence information from each paired sgRNA and predicts editing efficiency.
Software CRISPR eScreen is a sequence-sensitive model based on the Striped Hyena2 architecture that learns interpretable regulatory syntaxs and predicts activity. DeepDC is a deep learning model for predicting pgRNA activity on dual-cut fragment deletion by dual-SpCas9. Spatial and single cell eSpatial, a computational framework that deciphers spatially resolved enhancer regulation of gene expression by integrating spatial profiles of gene expression and chromatin accessibility. eMut, represents an integrative pipeline for detecting, imputing, and characterizing non-coding regulatory mutations with functional consequences at the single-cell resolution. eNet2.0, a comprehensive tool by introducing functions for network topology, comparative analysis, and dynamics, enabling systematic exploration of modular enhancer networks and their critical role in gene expression during development and disease. eNet1.0, an algorithm that constructs enhancer networks from single-cell multi-omics data to identify key genes and enhancers of cell identity and disease. Pesca, a computational method to predict and enhance spatial profiling of chromatin accessibility using scATAC-seq data. |